RNA-based treatments for select genetic diseases have made major headlines in the last few months by receiving FDA approval and giving hope to families of suffering children. They are Spinraza for spinal muscular atrophy (SMA) and Eteplirsen for Duchenne muscular dystrophy (DMD.)
These drugs are the future of therapeutics for genetic diseases - at the forefront of a long line of similarly based therapies coming down the pipeline. They are called "exon skipping" therapies and in order to understand how they work - we need to first understand what an exon is and why skipping it is an ingenious way to design a therapeutic.
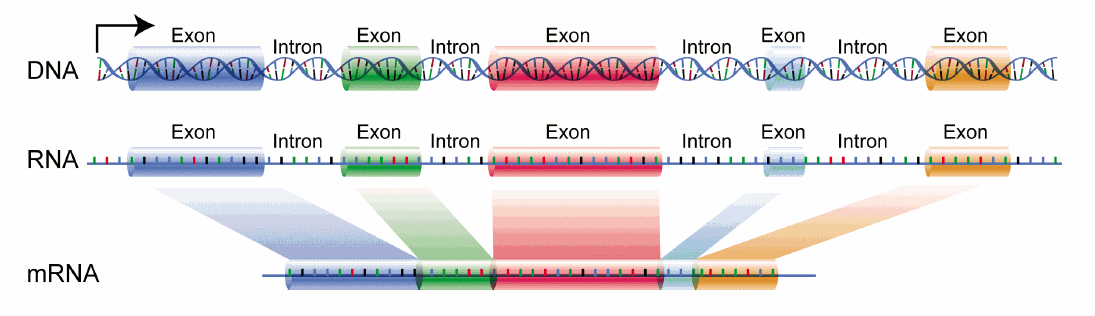
Last week, we wrote about how proteins are made from DNA entitled “The Central Dogma.” In it, we described how the piece of DNA that encodes a gene is made into a strand of RNA through the process of transcription. What we did not get into is that not all of the RNA that is transcribed off of DNA is used to make a protein. After a strand of RNA is made from the DNA, only portions of it (called exons - shown as the colored boxes in the figure below) will be used to make the protein. Those exons are stitched together through a process called splicing while the pieces of RNA that separate them, that will not be used, are cut out and removed (called introns.) The final mRNA, made up entirely of exons, will continue to be translated into protein.
Its kind’ve like when you take a piece of paper and cut out different parts all over it to make a snowflake. The small pieces on the edges get cut out and discarded (like introns) and what is left of the paper makes a beautiful snowflake (the protein.)
The process of splicing is complicated. One requirement are specific sequences on the RNA (called splice sites) to be available to the splicing machinery. If those splice sites are incorrect or blocked, the splicing process will be altered. When this happens, the final piece of mRNA will be different, and so will the protein that is made from it.
Certain diseases are caused by a protein that cannot function properly. A good example of this is the disease DMD. People who suffer with DMD have a mutation in the protein dystrophin. For some people with DMD, that mutation is located in exon number 51. (1)
And, manipulating the process of splicing to avoid that exon is the genius of exon skipping based therapies.
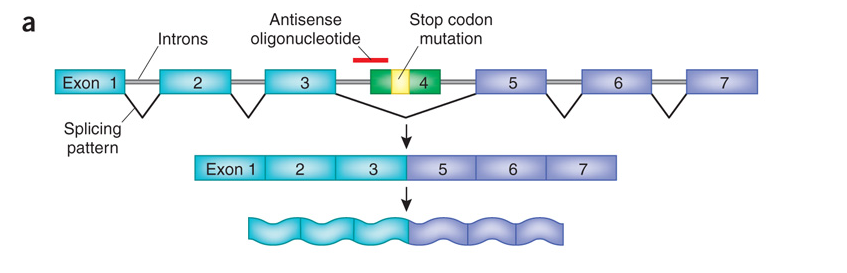
As shown in the figure below, a mutation is found in exon 4. The therapeutic is a small piece of RNA (the red line labeled 'antisense oligonucleotide') that lays over (and masks) the site that is required to splice exon 4. When the therapeutic is taken, exon 4 is essentially treated like an intron, skipped over, and left out of the final mRNA that creates the protein. The key here is that exon 4 is not required to make the protein. Although the protein that results may not be as good as in someone who never had the mutation in the first place - it is much better than having exon 4 left in the mRNA.
Eteplirsen and Spinraza are just the beginning of these drugs. Because muscular dystrophy is not always caused by a mutation in exon 51, Serepta, the company that designed Eteplirsen, has multiple other drugs in clinical development including a candidate for exon 53 skipping (SRP-4053) and a candidate for exon 45 skipping (SRP-4045.) They are working on other drugs to skip exons 44, 52, 50, 43, 55, 8 and 35.
Unfortunately, not all diseases can be treated through exon skipping. If skipping the exon creates a protein that does not function, or if the disease is caused by mutations in more than one gene, exon skipping may not be the answer.
But, for the thousands of families with children with DMD and SMA, exon skipping is a life saver. They are at the receiving end of years of hard work and innovation from the research labs of both academia and drug companies to see this basic biology be transformed into a disease cure.
Notes:
(1) Most proteins consist of roughly a dozen exons. Having 51 exons is highly unusual, but, Dystrophin is the largest protein in the cell.